🗺️ What You Will Learn in This Article
- Describe the double helix — base pairing rules, strand polarity, and the differences between A, B, and Z-DNA
- Compare prokaryotic vs eukaryotic DNA replication enzymes step by step
- Explain the roles of every key replication enzyme: helicase, primase, DNA pol I/III, ligase, topoisomerase
- Trace the leading and lagging strand synthesis including Okazaki fragments
- Identify all six major DNA repair pathways, their defective genes, and their associated cancer syndromes
- Connect telomere biology to ageing and cancer
- Answer any NEET PG / USMLE MCQ on DNA — recall, mechanism, or clinical scenario
📖 Introduction: Why This Topic Matters in Exams
A 35-year-old woman with no smoking history develops colon cancer. On further testing, her tumour shows microsatellite instability (MSI-High). Her mother and maternal aunt both had uterine cancer before age 50. The oncologist suspects a hereditary syndrome. This is Lynch syndrome — caused by a germline mutation in mismatch repair (MMR) genes. Without a clear understanding of DNA repair pathways, you would never connect a colon tumour’s molecular signature to a patient’s family history, let alone know which genes to test. This is exactly the kind of integrated question that now appears in NEET PG and USMLE Step 1.
DNA is one of the most consistently tested topics in PG entrance biochemistry. NEET PG and AIIMS ask it from multiple angles — structural details (Chargaff’s rules, base pair types), enzyme-level replication questions (which polymerase removes the primer?), and clinical MCQs connecting repair defects to specific cancer syndromes. USMLE Step 1 focuses heavily on the clinical genetics side: xeroderma pigmentosum, Lynch syndrome, Fanconi anaemia, and ataxia-telangiectasia all have disease-specific repair pathway defects that must be memorised precisely. FMGE tests the foundational structure and basic replication facts reliably every year.
This article covers DNA structure in full — from the double helix geometry to Z-DNA — then works through prokaryotic and eukaryotic replication enzyme by enzyme, followed by all six major repair pathways with their clinical disease links. High-yield tables, mnemonics, and five exam-standard MCQs follow at the end.
🔬 DNA Structure — From Nucleotide to Double Helix
The Nucleotide — Building Block of DNA
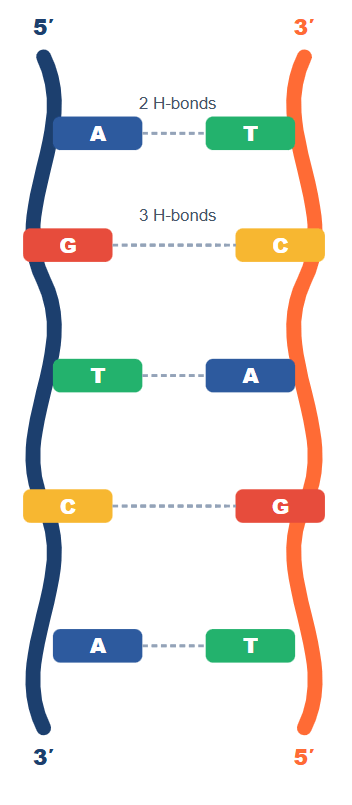
Each deoxyribonucleotide consists of three components: a deoxyribose sugar, a phosphate group, and a nitrogenous base. The bases are either purines (adenine [A] and guanine [G]) — two-ringed structures — or pyrimidines (cytosine [C] and thymine [T]) — single-ringed structures. In RNA, thymine is replaced by uracil (U).
The phosphodiester backbone runs 5′→3′ on each strand. This directionality is critical: all DNA polymerases synthesise only in the 5′→3′ direction. The two strands run antiparallel — one runs 5′→3′ while its complement runs 3′→5′.
Chargaff’s Rules — The Foundation of Base Pairing
Erwin Chargaff established the rules that made the double helix possible. In any double-stranded DNA: A pairs with T via 2 hydrogen bonds, and G pairs with C via 3 hydrogen bonds. This means the total amount of purines always equals pyrimidines. One practical consequence: the higher the G+C content of a DNA molecule, the higher its melting temperature (Tm), because G≡C bonds require more energy to break. This is tested frequently as “which organism’s DNA has the highest Tm?” — the answer is always the one with the highest G+C%.
A, B, and Z-DNA — Know the Differences
The table in Fig 1 summarises the three major DNA conformations. B-DNA is the physiological form — right-handed, 10 bp/turn, with a wide major groove where transcription factors bind. A-DNA forms under dehydrating conditions and also describes the conformation of RNA:DNA hybrids (important for understanding reverse transcription). Z-DNA is the only left-handed helix — it forms in regions rich in alternating purine-pyrimidine sequences, particularly at sites of active transcription. The “Z” name comes from its zigzag backbone appearance.
⚙️ DNA Replication — Enzymes, Steps & Mechanism
DNA replication is semiconservative (Meselson-Stahl experiment), bidirectional from origins of replication, and proceeds 5′→3′ only. Every enzyme in replication is a potential exam target. The table below is the highest-density study tool for this topic — learn every row.
| Enzyme | Function | Prokaryote | Eukaryote | Exam Hook |
|---|---|---|---|---|
| Helicase | Unwinds the double helix at the replication fork | DnaB | MCM complex | Requires ATP; moves 5′→3′ on lagging strand template |
| Primase | Synthesises short RNA primers (~10 nt) | DnaG | Pol α (with primase activity) | Only enzyme that can start a new strand de novo |
| DNA Pol III | Main replicative polymerase — high speed, high fidelity | ✅ Present | ❌ Absent | Has 3′→5′ exonuclease (proofreading); no 5′→3′ exo |
| DNA Pol I | Removes RNA primers (5′→3′ exonuclease) and fills gaps | ✅ Present | ❌ Absent | Only prokaryotic pol with 5′→3′ exonuclease activity |
| DNA Pol α | Initiates synthesis (with primase activity) | ❌ Absent | ✅ Present | Low fidelity; no proofreading |
| DNA Pol δ | Lagging strand synthesis; also DNA repair | ❌ Absent | ✅ Present | Has proofreading; works with PCNA (sliding clamp) |
| DNA Pol ε | Leading strand synthesis in eukaryotes | ❌ Absent | ✅ Present | High fidelity; associated with PCNA |
| DNA Ligase | Seals nicks — joins Okazaki fragments | NAD⁺ as cofactor | ATP as cofactor | Cofactor difference is a classic exam question |
| Topoisomerase I | Relaxes supercoils — cuts one strand | ✅ | ✅ | No ATP needed; target of camptothecin (anticancer) |
| Topoisomerase II | Relaxes supercoils — cuts both strands | Gyrase (prokaryotes) | ✅ | ATP needed; target of fluoroquinolones, etoposide |
| SSBs | Stabilise single-stranded DNA at fork | SSB proteins | RPA complex | Prevent re-annealing and secondary structure |
| Telomerase | Extends telomeres using built-in RNA template | ❌ Absent | ✅ (stem cells, cancer) | Reverse transcriptase; absent in somatic cells → ageing |
Leading vs Lagging Strand — The Core Concept
Because DNA polymerase can only synthesise 5′→3′, and the two template strands are antiparallel, replication cannot proceed the same way on both strands simultaneously.
- Leading strand — synthesised continuously 5′→3′ toward the replication fork. Requires one primer.
- Lagging strand — synthesised discontinuously, in short Okazaki fragments (~100–200 nt in eukaryotes, ~1000–2000 nt in prokaryotes), each starting with its own RNA primer. After synthesis, primers are removed by DNA Pol I (prokaryotes) / RNase H + FEN1 (eukaryotes), gaps are filled, and DNA ligase seals the nicks.
Telomeres and Telomerase
Telomeres are TTAGGG repeat sequences at the ends of linear eukaryotic chromosomes. They protect chromosomal ends from degradation and end-to-end fusion. Each round of replication shortens telomeres slightly because DNA polymerase cannot replicate the very end of the lagging strand template (the “end replication problem”).
Telomerase is a reverse transcriptase that carries its own RNA template (AAUCCC) and uses it to extend the 3′ end of the telomere, allowing subsequent priming and completion. It is highly active in germ cells, stem cells, and cancer cells. In most somatic cells it is absent — progressive telomere shortening acts as a molecular clock and tumour suppressor. Cancer cells reactivate telomerase to achieve replicative immortality (one of Hallmarks of Cancer).
🏥 DNA Repair Mechanisms — All Six Pathways with Clinical Links
This section is the most clinically important part of the topic. Each repair pathway is linked to a specific disease syndrome — and those syndromes appear in exam questions every year. The SVG below gives you the complete picture at a glance, followed by detailed descriptions of each pathway.
Nucleotide Excision Repair (NER) — Xeroderma Pigmentosum
NER handles bulky lesions that distort the double helix — primarily pyrimidine dimers (thymine-thymine dimers) caused by ultraviolet (UV) light, but also DNA adducts from chemicals like benzo[a]pyrene (cigarette smoke) and cisplatin (chemotherapy). The pathway recognises the distortion, cuts a ~27-nucleotide patch around the lesion on the damaged strand, and resynthesises using the intact strand as template. Seven XP proteins (XPA–XPG) carry out the recognition and incision steps.
A defect in any XP protein → Xeroderma Pigmentosum: extreme photosensitivity, >1000-fold elevated risk of UV-induced skin cancers (basal cell, squamous cell, melanoma), progressive neurodegeneration. Autosomal recessive. The key exam fact: these patients must avoid sunlight completely.
Mismatch Repair (MMR) — Lynch Syndrome
MMR corrects errors made during DNA replication — incorrect base insertions, deletions, and base-base mismatches that escape DNA Pol proofreading. The MutSα complex (MSH2/MSH6) recognises the mismatch; MutLα (MLH1/PMS2) coordinates excision and resynthesis. A key diagnostic signature: when MMR is defective, repetitive sequences called microsatellites accumulate insertions and deletions, producing microsatellite instability (MSI).
Lynch syndrome (HNPCC) is caused by germline mutations in MLH1, MSH2, MSH6, or PMS2. It is the most common hereditary colorectal cancer syndrome. Also significantly increases risk of uterine, ovarian, stomach, and urinary tract cancers. Amsterdam criteria and MSI-High tumour testing are key diagnostic tools.
Homologous Recombination (HR) — BRCA1, BRCA2, Fanconi Anaemia
HR repairs double-strand breaks (DSBs) using the sister chromatid as a template. It is error-free but only possible in S and G2 phases (when a sister chromatid is available). BRCA1 and BRCA2 are critical HR proteins — they facilitate RAD51-mediated strand invasion. Germline BRCA1/2 mutations confer dramatically elevated lifetime risk of breast (up to 85%) and ovarian (up to 44%) cancer. They also increase pancreatic and prostate cancer risk. BRCA-mutant tumours are exquisitely sensitive to PARP inhibitors (synthetic lethality).
Fanconi anaemia is caused by defects in genes of the FA pathway that coordinate with HR to repair interstrand crosslinks. Classic presentation: bone marrow failure, radial ray defects (absent thumbs), short stature, café-au-lait spots, predisposition to AML and squamous cell carcinomas. Chromosome breakage test with diepoxybutane (DEB) is diagnostic.
Non-Homologous End Joining (NHEJ) — Ataxia-Telangiectasia
NHEJ also repairs DSBs but does so by directly ligating the broken ends without a template — it is fast but error-prone. It operates primarily in G1 phase. It is the main mechanism for DSB repair in non-dividing cells and also mediates V(D)J recombination during lymphocyte development. ATM kinase is activated by DSBs and coordinates the cellular DNA damage response, including cell cycle arrest and apoptosis. ATM deficiency causes Ataxia-Telangiectasia: cerebellar ataxia, oculocutaneous telangiectasias, immunodeficiency, radiosensitivity, and very high risk of lymphoma and leukaemia.
🔄 How DNA Topics Connect to Other Subjects
- → Transcription & RNA Processing — DNA is the template for RNA synthesis; RNA Pol II reads the non-template strand 3′→5′ and synthesises mRNA 5′→3′. Promoter elements, enhancers, and transcription factors bind the major groove of B-DNA.
- → Translation & Genetic Code — the mRNA sequence derived from DNA determines the amino acid sequence of proteins via the triplet codon system. Mutations in DNA produce frameshift, missense, or nonsense changes in protein.
- → Cancer Biology — all six repair pathways, telomerase, and oncogenes/tumour suppressors link DNA biology to carcinogenesis. BRCA1/2, TP53, MLH1, and ATM are all tumour suppressor genes relevant here.
- → Pharmacology — Antibiotics — fluoroquinolones inhibit DNA gyrase (prokaryotic topoisomerase II). Novobiocin inhibits the B subunit of gyrase. These drugs exploit the difference between prokaryotic and eukaryotic topoisomerases.
- → Pharmacology — Anticancer Drugs — cisplatin creates crosslinks repaired by NER; PARP inhibitors exploit HR deficiency in BRCA mutant cells; topoisomerase inhibitors (camptothecin, etoposide) poison DNA repair machinery.
- → Genetics — Inheritance Patterns — Lynch, XP, Fanconi anaemia, and ataxia-telangiectasia are all autosomal recessive (except BRCA-associated cancers which follow autosomal dominant inheritance with loss of heterozygosity).
🎯 High-Yield Exam Facts
🔴 Z-DNA is the ONLY left-handed double helix
A and B are both right-handed. Z-DNA forms in GC-rich regions under negative supercoiling. Easily the most-asked structural fact in DNA conformation questions.
🔴 Prokaryotic DNA Pol I removes RNA primers (5′→3′ exonuclease); no eukaryotic polymerase has this activity
In eukaryotes, RNase H and Flap Endonuclease 1 (FEN1) remove primers instead. Confusing these is the single most common wrong answer in replication enzyme questions.
🔴 DNA Ligase cofactor: NAD⁺ in prokaryotes, ATP in eukaryotes
This one-liner is asked directly in exams. The antibiotic rifampicin does NOT target ligase — it targets RNA polymerase. Do not confuse.
🔴 NER defect → Xeroderma Pigmentosum; MMR defect → Lynch syndrome
These two repair-disease pairs are asked every single year. Know both directions: given a disease → name the pathway; given a defective protein → name the disease.
🟠 G≡C base pairs have 3 hydrogen bonds; A=T have 2
Higher G+C content = higher Tm (melting temperature). Thermophilic bacteria have high G+C content in their DNA for thermal stability.
🟠 Telomerase is a reverse transcriptase — it uses an RNA template to extend DNA
The RNA component is TERC (hTR), template sequence AAUCCC. The protein component (reverse transcriptase) is TERT. Active in cancer cells → target for anti-cancer drug development.
🟠 BRCA1/2 defects impair Homologous Recombination → sensitivity to PARP inhibitors
PARP inhibitors (olaparib, niraparib) cause synthetic lethality in BRCA-mutant cells. They are approved for BRCA-mutant breast, ovarian, and pancreatic cancers.
🟠 Ataxia-Telangiectasia = ATM kinase deficiency (NHEJ/DSB signalling)
Triad: cerebellar ataxia + oculocutaneous telangiectasias + immunodeficiency. Extremely radiosensitive — avoid unnecessary X-rays in these patients.
🟡 Okazaki fragments are longer in prokaryotes (~1000–2000 nt) than eukaryotes (~100–200 nt)
Commonly reversed in student answers. Eukaryotes have shorter fragments because of more frequent nucleosome positioning constraints.
🟡 Photolyase repairs pyrimidine dimers directly using visible light — but only in prokaryotes and lower eukaryotes, NOT in humans
Human NER (not photolyase) handles UV-induced pyrimidine dimers. A question about “photoreactivation” is asking about photolyase, not human NER.
🟡 Fanconi anaemia is diagnosed by chromosomal breakage test with diepoxybutane (DEB) or mitomycin C
DEB produces interstrand crosslinks that normal cells repair but Fanconi cells cannot — leading to excessive chromosomal breakage on karyotype.
🧠 Mnemonics & Memory Tricks
Mnemonic 1 — DNA Forms (A, B, Z)
“Always Be Careful with Z-Left”
- A-DNA → right-handed, dehydrated, RNA:DNA hybrids
- B-DNA → right-handed, physiological, wide major groove
- Z-DNA → left-handed (the only one!), GC-rich, zigzag backbone
💡 Exam trigger: Any question asking “which DNA form is left-handed?” — the answer is always Z.
Mnemonic 2 — DNA Repair Diseases
“Never Miss Hiding Broken DNA”
| Word | Pathway | Disease |
|---|---|---|
| Never | NER (Nucleotide Excision Repair) | Xeroderma Pigmentosum |
| Miss | MMR (Mismatch Repair) | Lynch Syndrome (HNPCC) |
| Hiding | HR (Homologous Recombination) | BRCA1/2 → Breast/Ovarian Ca; Fanconi Anaemia |
| Broken | BER (Base Excision Repair) | MYH-associated polyposis |
| DNA | DSB (NHEJ) | Ataxia-Telangiectasia (ATM) |
💡 Pro tip: Go through this mnemonic before any clinical genetics MCQ — it covers the five most testable repair-disease pairs in one pass.
Mnemonic 3 — Prokaryotic Replication Enzymes
“Pretty Girls Love Sexy Textbooks” (for the order of action)
- Primase (DnaG) — lays down RNA primer
- Gyrase / Topoisomerase — relieves supercoiling ahead of fork
- Leading strand — Pol III synthesises continuously
- SSBs — stabilise single-stranded template
- Trailer (lagging) strand — Pol III makes Okazaki fragments; Pol I removes primers; Ligase seals
💡 Use this to reconstruct the entire prokaryotic replication process from memory in an exam.
⚠️ Common Mistakes Students Make on This Topic
❌ Mistake: “Prokaryotes have DNA Pol I as their main replicative polymerase”
✅ Reality: DNA Pol III is the main replicative enzyme. DNA Pol I’s main role is primer removal (5′→3′ exonuclease) and gap filling.
📝 How it’s tested: “Which prokaryotic DNA polymerase removes RNA primers?” — answer is Pol I, not Pol III.
❌ Mistake: “XP (Xeroderma Pigmentosum) is caused by a base excision repair defect”
✅ Reality: XP is caused by a nucleotide excision repair (NER) defect. BER handles small, non-helix-distorting lesions; NER handles bulky adducts and pyrimidine dimers.
📝 How it’s tested: Clinical scenario of photosensitivity and skin cancer in a young child — the repair defect is NER, pathway, not BER.
❌ Mistake: “Telomerase works on the leading strand end problem”
✅ Reality: Telomerase extends the lagging strand template (the G-rich 3′ overhang of the parental strand) so that the lagging strand can be replicated. The “end problem” applies to the lagging strand, not the leading strand.
📝 How it’s tested: “Why do somatic cells shorten their telomeres with each division?” — answer: lagging strand end cannot be fully replicated; telomerase absent in somatic cells.
❌ Mistake: “All DNA polymerases have proofreading activity”
✅ Reality: DNA Pol α (which initiates synthesis in eukaryotes) has NO proofreading activity. This is why Pol δ and ε, which do have 3′→5′ exonuclease proofreading, take over after initiation.
📝 How it’s tested: “Which eukaryotic DNA polymerase lacks 3′→5′ exonuclease activity?” — Pol α.
❌ Mistake: “Lynch syndrome affects only the colon”
✅ Reality: Lynch syndrome (MMR defect) also significantly increases risk of endometrial (uterine), ovarian, stomach, small bowel, and urinary tract cancers. The classic exam trap is restricting it to colorectal cancer only.
📝 How it’s tested: “A family with colorectal cancer + endometrial cancer across multiple generations — which syndrome?” — Lynch syndrome.
📝 5 Practice MCQs — Test Yourself Now
Q1 — Pure Recall (Easy)
Which form of DNA is the ONLY left-handed double helix?
- A. A-DNA
- B. B-DNA
- C. Z-DNA
- D. H-DNA
👉 Click to reveal answer
Why correct: Z-DNA is the only left-handed double helix. It forms in GC-rich sequences under negative supercoiling and gets its name from the zigzag appearance of its phosphate backbone. Both A-DNA and B-DNA are right-handed.
Why A is wrong: A-DNA is right-handed, forms under dehydrating conditions, and is the conformation of RNA:DNA hybrid duplexes.
Why B is wrong: B-DNA is the standard physiological form — right-handed, 10 bp per turn, wide major groove.
Why D is wrong: H-DNA (triplex DNA) is a real structure but is not one of the standard three DNA conformations tested in PG exams.
Exam tip: The question “which DNA is left-handed?” is a guaranteed one-liner in FMGE and NEET PG. Z-DNA is the only correct answer every time.
Q2 — Mechanism (Easy–Medium)
In prokaryotic DNA replication, which enzyme is responsible for removing the RNA primer and filling in the gap with DNA?
- A. DNA Polymerase III
- B. DNA Polymerase I
- C. Primase
- D. DNA Ligase
👉 Click to reveal answer
Why correct: DNA Pol I is the ONLY prokaryotic polymerase with 5′→3′ exonuclease activity. This allows it to nick-translate — simultaneously removing the RNA primer ahead and filling in the gap with DNA. It is the essential primer removal enzyme in prokaryotes.
Why A is wrong: DNA Pol III is the main replicative enzyme but lacks 5′→3′ exonuclease activity — it cannot remove the primer. It only has 3′→5′ exonuclease (proofreading).
Why C is wrong: Primase (DnaG) synthesises the RNA primer — it does the opposite of what the question asks.
Why D is wrong: DNA Ligase seals the nick left after primer removal and gap filling — it acts after Pol I, not in the primer removal step itself.
Exam tip: Memorise: “Pol I removes, Pol III makes, Ligase seals.” This sequence describes the lagging strand completion in prokaryotes and appears as a direct question or embedded in a scenario about Okazaki fragment processing.
Q3 — Application (Medium)
A researcher blocks DNA topoisomerase II activity in a replicating bacterial cell. What is the most likely immediate consequence at the replication fork?
- A. Failure to remove RNA primers
- B. Accumulation of positive supercoils ahead of the fork, stalling replication
- C. Inability to seal Okazaki fragments
- D. Failure to unwind the double helix at the origin
👉 Click to reveal answer
Why correct: As helicase unwinds DNA at the fork, it introduces positive supercoiling ahead of the fork. DNA gyrase (prokaryotic Topoisomerase II) relieves this torsional stress by cutting both strands and re-joining them. If blocked, positive supercoils accumulate and physically prevent fork progression — replication stalls. This is the mechanism of action of fluoroquinolone antibiotics.
Why A is wrong: Primer removal is the job of DNA Pol I, not topoisomerase.
Why C is wrong: Sealing Okazaki fragments is the role of DNA Ligase.
Why D is wrong: Unwinding at the origin is the role of DnaA (initiator) and DnaB (helicase), not topoisomerase. Topoisomerase acts ahead of the already-unwinding fork.
Exam tip: Fluoroquinolones (ciprofloxacin, levofloxacin) target bacterial DNA gyrase (Topo II). Questions on fluoroquinolone mechanism always involve topoisomerase and supercoiling — never primer removal or strand sealing.
Q4 — Clinical Scenario (Medium–Hard)
A 28-year-old woman is referred to genetics after her brother is diagnosed with colon cancer at age 32. Her maternal grandmother had uterine cancer at 48 and a maternal uncle had gastric cancer at 55. Tumour testing of the brother’s cancer shows MSI-High status and loss of MLH1 expression on immunohistochemistry. Which of the following is the most likely diagnosis?
- A. Familial Adenomatous Polyposis (FAP)
- B. Lynch Syndrome (HNPCC)
- C. Xeroderma Pigmentosum
- D. BRCA2-associated colorectal cancer
👉 Click to reveal answer
Why correct: Lynch syndrome = early-onset colorectal cancer + family history of uterine/gastric/urinary cancers + MSI-High on tumour testing + loss of MMR protein (MLH1) on IHC. Every element of this vignette points to Lynch. It is caused by germline mutations in MMR genes (MLH1, MSH2, MSH6, PMS2).
Why A is wrong: FAP causes hundreds of colonic polyps (adenomatous), is caused by APC gene mutation, and does NOT cause MSI-High tumours. FAP tumours are microsatellite-stable.
Why C is wrong: XP (NER defect) causes UV-induced skin cancers and photosensitivity — it does not cause colorectal or uterine cancers.
Why D is wrong: BRCA2 mutations primarily increase risk of breast, ovarian, and pancreatic cancer. BRCA2-associated colorectal cancer is not a recognised clinical entity with MSI-High signature.
Exam tip: “MSI-High + young colorectal cancer + family history of uterine cancer” = Lynch syndrome every time. USMLE Step 1 loves this exact vignette pattern.
Q5 — Integration (Hard)
A child presents with progressive cerebellar ataxia, oculocutaneous telangiectasias, recurrent sinopulmonary infections, and markedly elevated alpha-fetoprotein (AFP). Karyotyping shows chromosomal instability. Which DNA repair pathway is defective and what is the specific protein involved?
- A. NER pathway — XPA protein
- B. MMR pathway — MLH1 protein
- C. DSB sensing/NHEJ — ATM kinase
- D. HR pathway — BRCA2 protein
👉 Click to reveal answer
Why correct: This is the classic presentation of Ataxia-Telangiectasia (A-T): cerebellar ataxia (progressive, beginning in childhood) + oculocutaneous telangiectasias + immunodeficiency (recurrent infections) + elevated AFP + chromosomal instability. The defective gene is ATM (Ataxia-Telangiectasia Mutated), a PI3K-like kinase that senses DSBs and orchestrates the DNA damage response including NHEJ coordination. These patients are exquisitely sensitive to ionising radiation.
Why A is wrong: XPA deficiency causes Xeroderma Pigmentosum — photosensitivity, skin cancers, NOT ataxia or telangiectasias.
Why B is wrong: MLH1 deficiency causes Lynch syndrome — colorectal/uterine cancers in adults, NOT childhood cerebellar ataxia.
Why D is wrong: BRCA2 deficiency causes breast/ovarian cancer predisposition in adults and Fanconi anaemia type D1 in childhood — the Fanconi presentation includes bone marrow failure and limb defects, not primarily ataxia and telangiectasias.
Exam tip: Elevated AFP in a child with ataxia is a high-yield distinguishing feature of A-T from other ataxia syndromes. It is asked in both NEET PG and USMLE as a one-liner and as a clinical vignette.
📚 References
- 📖 Harper’s Illustrated Biochemistry — Murray, Rodwell et al. | Chapter 35: DNA Organization, Replication & Repair
- 📖 Lippincott’s Illustrated Reviews: Biochemistry — Ferrier | Chapter 29: DNA Structure & Chapter 30: DNA Replication & Repair
- 📖 Biochemistry — Stryer, Berg, Tymoczko | Chapter 28: DNA Replication, Repair and Recombination
- 📖 Lehninger Principles of Biochemistry — Nelson & Cox | Chapter 25: DNA Metabolism
- 📖 Robbins Basic Pathology — Kumar, Abbas et al. | Chapter 7: Genetic and Pediatric Diseases (Repair Syndromes)
- 📖 Biochemistry for Medical Students — DM Vasudevan | Chapter on Molecular Biology
🚀 Keep Practising — You Are Not Done Yet
DNA, replication, and repair appear in 4–6 questions in most NEET PG papers. The theory you have just read is half the preparation — the other half is applying it cold under exam pressure. medicalmcq.in has hundreds of free Biochemistry MCQs with detailed explanations, all free.